Jun 23, 2020-Diverse bacteria and archaea rely on unusual methionine synthases. The Science…Methionine is one of the amino acids that make up proteins, and the final step in methionine synthesis is the transfer of a methyl group. Usually, the methyl group is obtained from methyl folates, but some anaerobic microbes use vitamin B12-binding proteins instead. By analyzing the genomes of diverse bacteria and archaea, we identified four families of folate-independent methionine synthases. Three of these families co-occur with B12-binding proteins, which indicates their likely partners. The fourth family does not require vitamin B12; instead, it obtains methyl groups from an oxygen-dependent partner protein.
Environmental Atlas
Sulfonate utilization by Desulfovibrio may expand ecological niches
September 18, 2019–Day, L.A.; K.B. De León, M.L. Kempher, J. Zhou, J.D. Wall (2019) Complete Genome Sequence of Desulfovibrio desulfuricans IC1, a Sulfonate-Respiring Anaerobe. Microbiology Resource Announcements. [doi]: 1128/MRA.00456-19
As part of ongoing work related to the 100-Well Survey of the Oak Ridge Reservation (ORR) supported by ENIGMA, Leslie Day, an undergraduate student, has published a genome announcement for a strain of Desulfovibrio, a sulfate-reducing bacterium (SRB), that grows by respiration of sulfonates. Genome comparisons have identified a candidate pathway and probes to explore the taxonomic distribution of this metabolism. Data from the earlier survey work showed that the SRB and sulfate were nearly mutually exclusive in the groundwater of the ORR prompting Leslie to determine if alternative sources of sulfate could possibly explain this result. A literature review pointed to sulfonates, ubiquitous in nature, as potential electron acceptors for SRB. As part of the Environmental Ark Campaign, she has shown that the sulfonate isethionate is used by a subset of SRB as a terminal electron acceptor and has identified the transporter and potential metabolic pathway for isethionate. The widespread distribution of sulfonates in the terrestrial environment may contribute to an understanding of the abundance of SRB in niches low in sulfate.
Energy Restriction Impacts Growth Physiology in a Sulfate-Reducing Biofilm
Krantz, G.P.; K. Lucas, E.L.-W. Majumder, L.T Hoang, R. Avci, G. Siuzdak and M.W. Fields (2019) Bulk phase resource ratio alters carbon steel corrosion rates and endogenously produced extracellular electron transfer mediators in a sulfate-reducing biofilm. Biofouling. [doi]:10.1080/08927014.2019.1646731 {PMID}:31402749 The presented results demonstrate that energy restriction (EAL v. EDL) greatly impacted biofilm growth and physiology […]
Plasmidome Retains Toxicity Tolerance ENIGMA publication highlighted by DOE
July 3, 2019–https://www.energy.gov/science/ber/articles/microbes-retain-toxicity-tolerance-after-they-escape-toxic-elements. * Kothari A, Wu Y-W, Charrier M, Rajeev L, Rocha AM, Paradis CJ, Hazen TC, Singer SW, Mukhopadhyay A. (2019) Large Circular Plasmids from Groundwater Plasmidomes Span Multiple Incompatibility Groups and Are Enriched in Multimetal Resistance Genes. mBio. [doi]:1128/mBio.02899-18
ENIGMA runs an internal proposal call to encourage early-career scientists to develop risky and interesting ideas that may contribute to our ongoing mission or changing direction by providing new data or tools. Here Ankita Kothari successfully managed a Discovery proposal from pitching the idea, developing the project to publication. This work was recently showcased at the national level by DOE
Bacteria often contain mobile genetic elements called plasmids that are circular DNA molecules that confer advantageous traits to the host and are capable of being horizontally transferred between different microbes. Studying the entire plasmid content (plasmidome) of a site can thus provide insights into genes that provide a selective advantage to a bacterial community. To study the incidence, diversity, and distribution of plasmids in groundwater along with exploring their role in environmental stress adaptation, we came up with the idea of exploring the plasmidome of the Oak Ridge Field Research Site. This Discovery proposal was risky because the existing plasmidome studies were limited to environments with high cell density. In contrast, at ENIGMA we focused on a challenging Department of Energy site characterized by low cell density and fluctuating populations. We developed and optimized methods to work with this environment and were successful in identifying over 600 different plasmids including several large plasmids that showed enrichment in metal tolerance genes in addition to antibiotic- and virus-resistance genes. Collaborations within ENIGMA helped to provide resources for environmental sampling, groundwater metal composition analysis, and sequence data analysis via KBase. This project now contributes to the 1) ENSuRC campaign since it continues to identify novel mobile genetic elements and accidentally discovered novel viruses along with 2) environmental ark campaign where the plasmids are being used to develop genetic tools to modify novel bacteria.
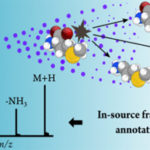
Adapting METLIN offers new metabolite identification opportunities
Domingo-Almenara, X.; J. R. Montenegro-Burke, C. Guijas, E. L.-W. Majumder, H. P. Benton and G. Siuzdak (2019) Autonomous METLIN-guided in-source fragmentation detection increases annotation confidence in untargeted metabolomics. ACS Analytical Chemistry. [doi]:1021/acs.analchem.8b03126 A clever adaptation of METLIN to turn an annotation problem into an identification advantage by Xavi Domingo and Scripps Research Institute team has […]
DuBSEQ functional characterization Nature Communications
January 18, 2019–ENIGMA researchers release DuBSEQ, a tool for broader use in microbial functional characterization and which allows for high throughput manipulation of specific strain libraries in Nature Communications.
A multi-disciplinary team of bench scientists and computational scientists collaborated to generate the large data set and analyses that confirm known functional genes and identify interesting genes of novel function.
Vivek K. Mutalik, Pavel S. Novichkov, Morgan N. Price, Trenton K. Owens, Sean Carim, Astrid Terry, Adam M. Deutschbauer, Adam P. Arkin
ENIGMA authors; Front row-Morgan N. Price, Pavel S. Novichkov, Vivek K. Mutalik, Astrid Terry, Sean Carim. Back row-Trenton K. Owens, Adam M. Deutschbauer, Adam P. Arkin — In Emeryville, California, 1/16/2019
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria https://doi.org/10.1038/s41467-018-08177-8