The Genomic Features Underlying Adaptation of Rhodanobacter Strains to Stressful Conditions (denitrification processes, heavy metal resistance, and restriction-modification systems) present at ENIGMA Field Site ORR. Rhodanobacter species have been found dominant in the subsurface of Oak Ridge Reservation (ORR) site contaminated with acid, nitrate, metal radionuclides, and other heavy metals. Recently, multiple Rhodanobacter strains have […]
Environmental Atlas
Sampling Down Well In Situ Microbial Colonization Conditions
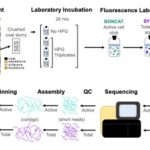
LJ McKay*, HJ Smith*, E Barnhart, HD Schweitzer, RR Malmstrom, D Goudeau, MW Fields (2021) Activity-based, genome-resolved metagenomics uncovers key populations and pathways involved in subsurface conversions of coal to methane Nature ISME Journal [DOI]:10.1038/s41396-021-01139-x PMID: 34689183 Development of activity-based metagenomics, enabled the identification of translationally active microorganisms and associated functional capacity Demonstration on terrestrial subsurface […]
A Simple, Cost-Effective, and Automation-Friendly Direct PCR Approach for Bacterial Community Analysis
November 22, 2021 Simple, cost-effective, and automation-friendly direct-PCR-based 16S rRNA sequencing method to study the dynamics, microbial interaction, and assembly of various microbial communities in a high-throughput format
Host Bacteria Are Helped By Viral Genomes
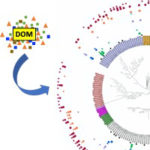
Kothari, Ankita.; S. Roux, H. Zhang, A. Prieto, J-M Chandonia, S. Spencer, X. Wu, A M. Deutschbauer, A. P. Arkin, E J. Alm, R. Chakraborty, A Mukhopadhyay (2021) Ecogenomics of groundwater viruses suggests niche differentiation linked to specific environmental tolerance. mSystems [doi]:1128/mSystems.00537-21 {PMID}: 34184913 PMCID: PMC8269241 OSTI:1807913 Niche Specific Genes from Phage Genomes are Advantageous for Communities […]
Understanding how microbes adapt to stressful conditions
Adaptation to previous stressors influence which genes acquire mutations during experimental evolution M.L. Kempher, X. Tao, R. Song, B. Wu, D.A. Stahl, J.D. Wall, A.P. Arkin, A. Zhou, J. Zhou. “Effects of genetic and physiological divergence on the evolution of a sulfate-reducing bacterium under conditions of elevated temperature.” mBio, 11(4), e00569-20 (2020). [doi: 10.1128/mBio.00569-20] The […]
Culturing of “Unculturable” Subsurface Microbes: Natural Organic Carbon Source Fuels the Growth of Diverse and Distinct Bacteria from Groundwater
June 8 2021. An effective cultivation and isolation strategy using natural complex carbon source recovers diverse and novel bacteria from the terrestrial subsurface. The Science…. Despite rapid technological advances in modern molecular tools for identification of key microbial taxa and critical metabolic processes in a given environment, a complete interpretation of omics-based data is still constrained by the unavailability of reference genomes and physiologically characterized isolates. Challenges in microbial cultivation/isolation in the laboratory have impeded the ability of microbiologists to fully investigate the roles and function of microbes in terrestrial subsurface ecosystems. To date scientists have only been able to cultivate less than 2% of microbes on Earth in the laboratory.
New Tools Allow for Faster Mapping of Large Numbers of Phage Resistant Genes
Oct 13, 2020-
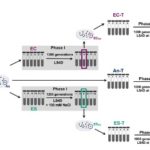
Mutalik, V.K.; B.A. Adler, H.S. Rishi, D. Piya, C. Zhong, B. Koskella, E.M. Kutter, R. Calendar, P.S. Novichkov, M.N. Price, A.M. Deutschbauer, A.P. Arkin (2020) High-throughput mapping of the phage resistance landscape in E. coli. PLoS Biology [doi]:10.1371/journal.pbio.3000877 OSTI:1714384
Phage work supported by ENIGMA Discovery to develop new tools for use towards mission critical questions is highlighted in https://newscenter.lbl.gov/2020/12/10/phages-natures-hidden-arsenal/
EISA-MRM: Quantitative MS Toward A $5 Metabolome
August 24, 2020-Xue J, Guijas C, Benton HP, Warth B, Siuzdak G (2020) METLIN MS2 molecular standards database: a broad chemical and biological resource. Nature Methods. 17(10) 953-954 [doi]: 10.1038/s41592-020-0942-5, PMID: 32839599
SCIENCE: A key ENIGMA Aim is linking field data to the phenotypic characterization of isolates, testing mechanistic hypothesis in the laboratory.
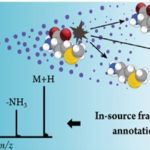
EISA-MRM (Enhanced In-Source Fragmentation/Annotation Multiple Reaction Monitoring):
This novel quantitative mass spectrometry technique allows the measurement of concentrations of the metabolites in ENIGMA samples using much cheaper instrumentation. These efforts characterizing key metabolites developing experimental quantitative methods to monitor the change of metabolome in bacteria are mission critical. The techniques will be used to design and test field-derived hypothesis in the laboratory.