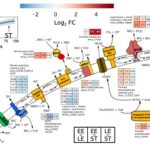
Shi, Z.; H. Yin, J.D. Van Nostrand, J.W. Voordeckers, Q. Tu, Y. Deng, M. Yuan, A. Zhou, P. Zhang, N. Xiao, D. Ning, Z. He, L. Wu and J. Zhou (2019) Functional Gene Array-Based Ultrasensitive and Quantitative Detection of Microbial Populations in Complex Communities. International Society for Microbial Ecology Journal. [doi]:10.1128/mSystems.00296-19 {PMID}:31213523PMCID:PMC6581690 OSTI:1580884 The rapid development […]
ENIGMA
Plasmidome Retains Toxicity Tolerance ENIGMA publication highlighted by DOE
July 3, 2019–https://www.energy.gov/science/ber/articles/microbes-retain-toxicity-tolerance-after-they-escape-toxic-elements. * Kothari A, Wu Y-W, Charrier M, Rajeev L, Rocha AM, Paradis CJ, Hazen TC, Singer SW, Mukhopadhyay A. (2019) Large Circular Plasmids from Groundwater Plasmidomes Span Multiple Incompatibility Groups and Are Enriched in Multimetal Resistance Genes. mBio. [doi]:1128/mBio.02899-18
ENIGMA runs an internal proposal call to encourage early-career scientists to develop risky and interesting ideas that may contribute to our ongoing mission or changing direction by providing new data or tools. Here Ankita Kothari successfully managed a Discovery proposal from pitching the idea, developing the project to publication. This work was recently showcased at the national level by DOE
Bacteria often contain mobile genetic elements called plasmids that are circular DNA molecules that confer advantageous traits to the host and are capable of being horizontally transferred between different microbes. Studying the entire plasmid content (plasmidome) of a site can thus provide insights into genes that provide a selective advantage to a bacterial community. To study the incidence, diversity, and distribution of plasmids in groundwater along with exploring their role in environmental stress adaptation, we came up with the idea of exploring the plasmidome of the Oak Ridge Field Research Site. This Discovery proposal was risky because the existing plasmidome studies were limited to environments with high cell density. In contrast, at ENIGMA we focused on a challenging Department of Energy site characterized by low cell density and fluctuating populations. We developed and optimized methods to work with this environment and were successful in identifying over 600 different plasmids including several large plasmids that showed enrichment in metal tolerance genes in addition to antibiotic- and virus-resistance genes. Collaborations within ENIGMA helped to provide resources for environmental sampling, groundwater metal composition analysis, and sequence data analysis via KBase. This project now contributes to the 1) ENSuRC campaign since it continues to identify novel mobile genetic elements and accidentally discovered novel viruses along with 2) environmental ark campaign where the plasmids are being used to develop genetic tools to modify novel bacteria.
ENIGMA teamwork is featured in Elements
June 24, 2019–A highly collaborative Discovery project combines two techniques BONCAT+FACS, (Bioorthogonal Non-Canonical Amino Acid Tagging, and Fluorescent Activated Cell Sorting), to detect individual active microbes in soil from our Oak Ridge Field Research Site in Tennessee. To find out more https://newscenter.lbl.gov/2019/06/24/new-microbial-research-technique/
Couradeau E.; J. Sasse, D. Goudeau, N. Nath, R. Chakraborty, T.C. Hazen, B.. Bowen, R.R. Malmstrom, T.R. Northen (2019) Challenging our view of the active fraction of soil microbiomes using BONCAT-FACS-Seq. Nature Communication. [doi]:1038/s41467-019-10542-0. Estelle Couradeau – 2018 ENIGMA Strategic Planning – Lawrence Berkeley National Laboratory on Tuesday, October 9, 2018 in Berkeley, Calif. 10/09/18
Adapting METLIN offers new metabolite identification opportunities
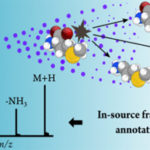
Domingo-Almenara, X.; J. R. Montenegro-Burke, C. Guijas, E. L.-W. Majumder, H. P. Benton and G. Siuzdak (2019) Autonomous METLIN-guided in-source fragmentation detection increases annotation confidence in untargeted metabolomics. ACS Analytical Chemistry. [doi]:1021/acs.analchem.8b03126 A clever adaptation of METLIN to turn an annotation problem into an identification advantage by Xavi Domingo and Scripps Research Institute team has […]
Challenging Carbon Nitrogen Ratios in a Dual-Pathway Actinobacterium
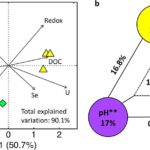
March 3, 2021-Vuono D.C., Read R.W., Hemp J., Sullivan B.W., Arnone, J.A., Neveux I., Blank R.R., Loney E., Miceli D., Winkler M-K.H., Chakraborty R., Stahl D.A., Grzymski J.J. 2019. Resource Concentration Modulates the Fate of Dissimilated Nitrogen in a Dual-Pathway Actinobacterium. Frontiers in Microbiology Vol 10, 3 [doi]:3389/fmicb.2019.00003{PMID}:30723459 PMCID:PMC6349771 OSTI:156058. In this study, we tested the effects of C:NO3− ratio and substrate concentration on pathway selection between ammonification and denitrification in the dual-pathway organism Intrasporangium calvum C5. We challenge the paradigm that C:NO3− ratio controls pathway selection based on a simple principle: ratios do not account for substrate concentrations, which can impose resource limitation for C or NO3−
DuBSEQ functional characterization Nature Communications
January 18, 2019–ENIGMA researchers release DuBSEQ, a tool for broader use in microbial functional characterization and which allows for high throughput manipulation of specific strain libraries in Nature Communications.
A multi-disciplinary team of bench scientists and computational scientists collaborated to generate the large data set and analyses that confirm known functional genes and identify interesting genes of novel function.
Vivek K. Mutalik, Pavel S. Novichkov, Morgan N. Price, Trenton K. Owens, Sean Carim, Astrid Terry, Adam M. Deutschbauer, Adam P. Arkin
ENIGMA authors; Front row-Morgan N. Price, Pavel S. Novichkov, Vivek K. Mutalik, Astrid Terry, Sean Carim. Back row-Trenton K. Owens, Adam M. Deutschbauer, Adam P. Arkin — In Emeryville, California, 1/16/2019
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria https://doi.org/10.1038/s41467-018-08177-8
Using High-Throughput Genetics to Discover a Novel Catabolic Pathway
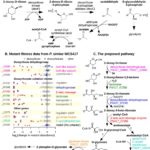
January, 12 2019-Price, M. N.; J. Ray, A. T. Iavarone, H. K. Carlson, E. M. Ryan, R. R. Malmstrom, A. P. Arkin, A. M. Deutschbauer (2019) An Oxidative Pathway of Deoxyribose Catabolism. mSystems. [doi]:1128/mSystems.00297-18
As part of the Environmental Ark campaign to characterize bacteria found at our Oak Ridge Research Field Site in Tennessee, highly collaborative work between ENIGMA and JGI generates data from sediment to use high-throughput genetics to discover a novel catabolic pathway as the first of many. Diverse bacteria use oxidative pathways to consume deoxyribonate (a waste product of metabolism).
David Stahl Honored by Seattle Aquarium
January 8, 2019–We know David Stahl as a Principal Investigator in ENIGMA working in part on the microbial conversion of ammonia to nitrate, a major step in the global nitrogen cycle.
Not only is he an internationally renowned microbial ecologist, elected to the National Academy of Engineering in 2012; now he is also a local Seattle celebrity! His discovery, that ammonia-oxidizing archaea (AOA) microorganisms are the dominant nitrifiers in most natural systems, was honored by a plaque at the Seattle Aquarium. His team was first to isolate AOA microorganisms from a water sample taken from a fish tank at the Seattle Aquarium.
https://www.ce.washington.edu/news/article/2019-01-03/david-stahls-discovery-honored-seattle-aquarium