Basics2Breakthroughs: Observing the microbial genome from Berkeley Lab on YouTube. ENIGMA Project Scientist Lauren Lui is profiled on LBL Basics2Breakthroughs, filmed for LBNL’s 90th anniversary. She uses the latest bioinformatics technology to study microbes so we can learn how their genes shape the environment. Her ultimate goal is to be able to predict microbial interactions […]
News Articles Archives
Environmental exposure history is critical in groundwater contaminant attenuation
Paradis, CJ, JI Miller, J-W Moon, SJ Spencer, LM Lui, JD Van Nostrand, D Ning, AD Steen, LD McKay, AP Arkin, J Zhou, EJ Alm, TC Hazen Sustained Ability of a Natural Microbial Community to Remove Nitrate from Groundwater (2021) Groundwater [DOI]:10.1111/gwat.13132 OSTI:1807518 A consortium of ENIGMA researchers at University of Tennessee, Oak Ridge National […]
A Simple, Cost-Effective, and Automation-Friendly Direct PCR Approach for Bacterial Community Analysis
November 22, 2021 Simple, cost-effective, and automation-friendly direct-PCR-based 16S rRNA sequencing method to study the dynamics, microbial interaction, and assembly of various microbial communities in a high-throughput format
Automating Culturing Technology Aids In Scaling Bacterial Growth Experiments
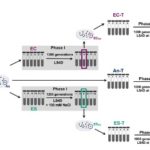
An automated multiplexed turbidometric and data collection system for measuring growth kinetics of anaerobes dependent on gaseous substrates Hunt, KA; J Forbes, F Taub, N Elliott, J Hardwicke, R Petersen, N Stopnisek, DAC Beck, DA Stahl; Journal of Microbiological Methods, [DOI]:10.1016/j.mimet.2021.106294 For decades, the cloudiness (turbidity) of a culture has been used to monitor the […]
Fostering interest in science and engineering through community college outreach
Recently, ENIGMA Principal Investigator Romy Chakraborty hosted college student Mariam Alsaid; an intern who participated in the The Transfer-to-Excellence Research Experiences for Undergraduates (TTE REU)program. She worked with ENIGMA postdocs Mon Oo Yee, and Kristine Cabugao. Mariam shared her experience with the program in the following interview
Host Bacteria Are Helped By Viral Genomes
Kothari, Ankita.; S. Roux, H. Zhang, A. Prieto, J-M Chandonia, S. Spencer, X. Wu, A M. Deutschbauer, A. P. Arkin, E J. Alm, R. Chakraborty, A Mukhopadhyay (2021) Ecogenomics of groundwater viruses suggests niche differentiation linked to specific environmental tolerance. mSystems [doi]:1128/mSystems.00537-21 {PMID}: 34184913 PMCID: PMC8269241 OSTI:1807913 Niche Specific Genes from Phage Genomes are Advantageous for Communities […]
Understanding how microbes adapt to stressful conditions
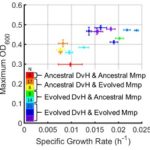
Adaptation to previous stressors influence which genes acquire mutations during experimental evolution M.L. Kempher, X. Tao, R. Song, B. Wu, D.A. Stahl, J.D. Wall, A.P. Arkin, A. Zhou, J. Zhou. “Effects of genetic and physiological divergence on the evolution of a sulfate-reducing bacterium under conditions of elevated temperature.” mBio, 11(4), e00569-20 (2020). [doi: 10.1128/mBio.00569-20] The […]
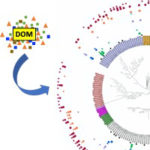
Culturing of “Unculturable” Subsurface Microbes: Natural Organic Carbon Source Fuels the Growth of Diverse and Distinct Bacteria from Groundwater
June 8 2021. An effective cultivation and isolation strategy using natural complex carbon source recovers diverse and novel bacteria from the terrestrial subsurface. The Science…. Despite rapid technological advances in modern molecular tools for identification of key microbial taxa and critical metabolic processes in a given environment, a complete interpretation of omics-based data is still constrained by the unavailability of reference genomes and physiologically characterized isolates. Challenges in microbial cultivation/isolation in the laboratory have impeded the ability of microbiologists to fully investigate the roles and function of microbes in terrestrial subsurface ecosystems. To date scientists have only been able to cultivate less than 2% of microbes on Earth in the laboratory.