The Science
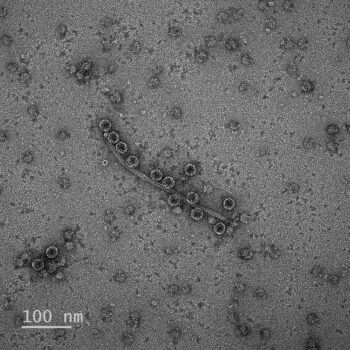
Bacteriophages, or phages, are viruses that target bacteria. Some of the smallest bacteriophages on earth use single-stranded RNA as their genetic material. While multiple genes are involved in programmed host lysis for double-stranded DNA phages, small single-stranded RNA phages require only a single gene. However, the mode of action of these lysis systems is poorly understood. In this study, published in Nature Chemical Biology, ENIGMA researchers used a newly-developed high-throughput genetic screen to uncover all potential proteins targeted by a diverse suite of single gene lysis systems. They found that single-gene lysis systems, despite belonging to different phages, may converge towards a common and conserved targeting mechanism.
The Impact
With new, high-throughput genomic technologies, researchers can now entirely sequence the RNA material from environmental samples, including those from soil and water. With this new capability, researchers are discovering that small, single-stranded RNA phages that infect bacteria are highly abundant in the environment, but it remains unknown which bacteria they infect and how they kill target bacteria. In this study, ENIGMA researchers present their newly-developed approach for investigating how small single gene lysis systems encoded by RNA phages can harm target bacteria. This approach could be broadly applied to study the mode of action for diverse phage-encoded systems as well as toxic bacterial genes.
Summary
Despite the importance of phages in microbial ecology, evolution, virulence and fitness, our knowledge of the gene functions encoded by phages is limited to a handful of well-studied double stranded DNA (dsDNA) phages. Recent metatranscriptomic studies have revealed that the true ubiquity and diversity of non-dsDNA phages, including single-stranded RNA-phages (ssRNA-phages), has been largely underestimated, and it is imperative to develop new methods to investigate and harness their genetic diversity. In this study, researchers develop a new method to characterize single gene encoded lysis systems of single strand RNA phages. These lytic proteins are also known as “protein antibiotics,” as they share mechanistic similarity in their action to the majority of antibiotic classes. Using the new barcoded screening technology, researchers uncovered genome-wide targets of diverse ssRNA-phage-derived protein antibiotics to efficiently identify the host mechanisms targeted in bacteriolysis. Their high-throughput suppressor screen revealed the importance of genes involved in cell-wall biosynthesis as targets and mediators of diverse single-gene lysis (Sgl) activity. Specifically, they uncovered the the protein Lipid II flippase, MurJ, as the target for Sgl-PP7 and Sgl-M. These two unrelated Sgl proteins belong to two different phages and infect different hosts, suggesting a case of convergent evolution and the potential presence of universal “weak spots” in the cell wall. The novel approach could have significant applications for characterizing the hundreds of putative Sgl identified in genomic databases and recent studies.
Contact

Vivek K Mutalik (vkmutalik@lbl.gov) and Adam Arkin (aparkin@lbl.gov)
Lawrence Berkeley National Laboratory
Adler, B.A.; K. Chamakura, H. Carion, J. Krog, A.M. Deutschbauer, R. Young, V.K. Mutalik and A.P. Arkin (2023) Multicopy suppressor screens reveal convergent evolution of single-gene lysis proteins. Nature Chemical Biology [DOI]:1038/s41589-023-01269-7 OSTI:1960236
Funding
The Microbiology Program of the Innovative Genomics Institute, Berkeley, funded this project. The technology work was funded by ENIGMA (Ecosystems and Networks Integrated with Genes and Molecular Assemblies), a Science Focus Area Program at Lawrence Berkeley National Laboratory, is based on work supported by the Department of Energy Office of Science, Office of Biological and Environmental Research.
Related Links
A Quick New Way to Screen Virus Proteins for Antibiotic Properties, Lawrence Berkeley National Laboratory News, Berkeley, CA
Uncovering the Secrets of the Smallest Phages Innovative Genomics Institute News, Berkeley, CA
Dub-seq Used to Screen Phage Proteins for Antibiotic Properties, Biosciences News, Lawrence Berkeley National Laboratory, Berkeley, CA
Dual-Barcoded Shotgun Expression Library Sequencing (Dub-seq) Intellectual Property Office (IPO), Lawrence Berkeley National Laboratory News, Berkeley, CA
