 Lawrence Berkeley National Lab
Lawrence Berkeley National Lab
Environmental Genomics & Systems Biology
SFA Technical Co-Manager
aparkin@lbl.gov
(510) 643-5678
Adam Arkin is an expert in the systems and synthetic biology of microbes. His role as the SFA Technical Co-Manager is to ensure the scientific basis of the mission goals, working with the Science Leads and Principal Investigators to develop coherent projects that address high-value problems, especially those of scale. He helps Principal Investigators and Project Scientists effectively plan and execute experiments addressing relevant themes.
The Arkin lab is specifically engaged in developing the metafunctional genomic methods to identify the most important microbes and their environmental correlates in the field to drive prioritizations for isolation and enrichment in the lab; developing new genetic, culturing, and computational methods for mapping genotype-to-phenotype in these laboratory systems; and developing the integrated mechanistic microbial modeling systems for predicting community dynamics under perturbations in the field and laboratory.
Relevant Publications
 Huang Explores How Bacteria Store Carbon at 2024 BioEPIC Research SLAM - Congratulations to Joshua Jiaqi Huang, a UC Berkeley graduate student researcher in Biosciences senior faculty scientist Adam Arkin’s group, who represented ENIGMA and Biosciences at the 2024 BioEPIC Research SLAM. More →
Huang Explores How Bacteria Store Carbon at 2024 BioEPIC Research SLAM - Congratulations to Joshua Jiaqi Huang, a UC Berkeley graduate student researcher in Biosciences senior faculty scientist Adam Arkin’s group, who represented ENIGMA and Biosciences at the 2024 BioEPIC Research SLAM. More →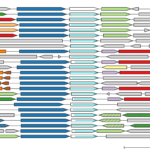 fast.genomics: A Fast Comparative Genome Browser for Diverse Bacteria and Archaea - ENIGMA researchers have developed fast.genomics, a new tool which allows users to search for a protein and identify other proteins that share a common ancestor or are located in similar genomic regions. More →
fast.genomics: A Fast Comparative Genome Browser for Diverse Bacteria and Archaea - ENIGMA researchers have developed fast.genomics, a new tool which allows users to search for a protein and identify other proteins that share a common ancestor or are located in similar genomic regions. More → Multi-disciplinary research develops a deeper understanding of subsurface microbiology - ENIGMA is organized into three aims that reflect the different research scales (i.e., field level to molecular level) required to build a comprehensive picture of subsurface microbiology at the Oak Ridge Reservation (ORR): Subsurface Observatory, Environmental Atlas, and Environmental Simulations. Several key research initiatives, described in this cross-aim overview, highlight this framework in action. More →
Multi-disciplinary research develops a deeper understanding of subsurface microbiology - ENIGMA is organized into three aims that reflect the different research scales (i.e., field level to molecular level) required to build a comprehensive picture of subsurface microbiology at the Oak Ridge Reservation (ORR): Subsurface Observatory, Environmental Atlas, and Environmental Simulations. Several key research initiatives, described in this cross-aim overview, highlight this framework in action. More →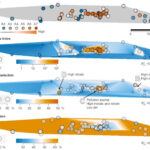 Nuclear Waste Sites Yield Microbial Ecosystem Insights - In a flagship seven-year study, published this January in the journal Nature Microbiology, ENIGMA researchers explored how environmental stresses influence the composition and structure of microbial communities in the groundwater of the Oak Ridge Reservation (ORR), a former nuclear waste disposal site. More →
Nuclear Waste Sites Yield Microbial Ecosystem Insights - In a flagship seven-year study, published this January in the journal Nature Microbiology, ENIGMA researchers explored how environmental stresses influence the composition and structure of microbial communities in the groundwater of the Oak Ridge Reservation (ORR), a former nuclear waste disposal site. More → ENIGMA researchers Win 2nd Annual BioEPIC SLAM - Adam Arkin, lead scientist, along with ENIGMA team members Bradley Biggs , a postdoctoral researcher in the Arkin lab, and Mingfei Chen, a postdoctoral researcher in the Chakraborty lab, recently participated in a virtual Research Slam promoting science related to the BioEPIC building. https://bioepic.lbl.gov/slam/ Results: Mingfei Chen (ENIGMA) took first place, Mo Ombadi (Watershed Function) second, and Bradley Biggs… More →
ENIGMA researchers Win 2nd Annual BioEPIC SLAM - Adam Arkin, lead scientist, along with ENIGMA team members Bradley Biggs , a postdoctoral researcher in the Arkin lab, and Mingfei Chen, a postdoctoral researcher in the Chakraborty lab, recently participated in a virtual Research Slam promoting science related to the BioEPIC building. https://bioepic.lbl.gov/slam/ Results: Mingfei Chen (ENIGMA) took first place, Mo Ombadi (Watershed Function) second, and Bradley Biggs… More → Examining adaptations of a dominant bacterial species at a contaminated subsurface site - Leveraging comparative genomics, community analysis, and geochemical data to understand adaptations that facilitate survival of a dominant community member in the contaminated Oak Ridge Reservation subsurface. More →
Examining adaptations of a dominant bacterial species at a contaminated subsurface site - Leveraging comparative genomics, community analysis, and geochemical data to understand adaptations that facilitate survival of a dominant community member in the contaminated Oak Ridge Reservation subsurface. More →