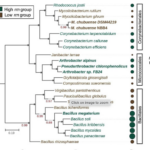
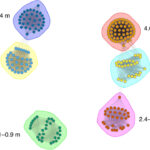
Microorganisms have evolved various life-history strategies to survive fluctuating resource conditions in soils. However, it remains elusive how the life-history strategies of microorganisms influence their processing of organic carbon, which may affect microbial interactions and carbon cycling in soils. Here, ENIGMA researchers characterized the genomic traits, exometabolite profiles, and interactions of soil bacteria representing copiotrophic and oligotrophic strategists. I
ENIGMA
Two model phages characterized by new CRISPR based technology extensible to diverse phages
ENIGMA researchers demonstrated for the first time that they can, on a genome-wide scale, identify phage genes that are essential (or not) to infecting bacteria, and then replace non-essential DNA with distinctive barcode tags. Their method could unlock potent biotechnology applications.
Mixed Nitrate and Metal Contamination Influences Operational Speciation of Toxic and Essential Elements
ENIGMA researchers compared the availability of elements in non-contaminated and acidic nitrate contaminated sediment and found that contamination impacted the bioavailability of essential elements.
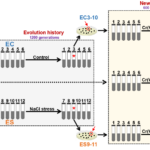
Environmental bacteria may have candidate genes for bioremediation
ENIGMA researchers identified important genes for improving Chromate resistance in genetically diverse bacteria through experimental evolution.
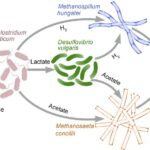
To Study Competition and Cross-Feeding, Scientists Build Synthetic Microbiomes
A new study investigated complex interactions among four cross-feeding microorganisms in a synthetic community (SynCom) that converts cellulose to methane and carbon dioxide, paving the way toward a more predictive understanding of the impact of environmental perturbations on microbial interactions sustaining geochemically significant processes in natural systems.
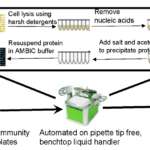
Novel Protocol Leverages Automatic Liquid Transfer to Prepare Hundreds of Microbial Cell Cultures for Proteomic Analysis
ENIGMA researchers detailed a step-by-step protocol that consists of cell lysis in alkaline chemical buffer (NaOH/SDS) followed by protein precipitation with high-ionic strength acetone in 96-well format. This protocol, combined with previously established automated protein quantification and protein normalization protocols, provides a rapid, cost-effective method to prepare LC-MS proteomic samples from bacteria and non-filamentous fungi cell cultures.
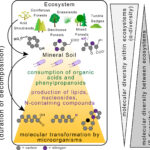
Decomposition Decreases Molecular Diversity and Ecosystem Similarity of Soil Organic Matter
ENIGMA researchers collaborated with the Lehmann Lab at Cornell University to find that microbial decomposition drives significant variability in the molecular richness and diversity of soil organic matter between soil horizons and ecosystems.
Microbial Communities Vary Widely Underground, Even When Close
Study finds that geochemistry and hydrogeology can influence community type and function, even when those communities are only 10 cm apart.