ENIGMA is organized into three aims that reflect the different research scales (i.e., field level to molecular level) required to build a comprehensive picture of subsurface microbiology at the Oak Ridge Reservation (ORR): Subsurface Observatory, Environmental Atlas, and Environmental Simulations. Several key research initiatives, described in this cross-aim overview, highlight this framework in action.
Rainfall Impacts Hydrology Of Test Site More Than Field Sampling
ENIGMA researchers measured physical and geochemical effects of cone penetrometer testing on nearby shallow groundwater wells. They found that the effects of cone penetration on water level and conductivity were minimal.
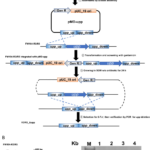
Genome Editing in Rhodanobacter denitrificans
ENIGMA researchers demonstrated the development and application of a markerless deletion mutagenesis system in nitrate-reducing bacterium Rhodanobacter denitrificans. This method marks a crucial step in advancing Rhodanobacter as a model denitrifying bacterium for the study of denitrification in groundwater ecosystems and diverse molecular mechanisms of low-pH resistance.
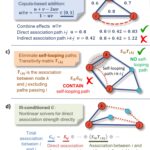
Disentangling Direct and Indirect Relationships in Association Networks
New iDIRECT framework could help scientists better understand biological systems by disentangling direct from indirect relationships in association networks.
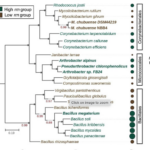
Bacterial community composition impacts the transformation of carbon in soils
Microorganisms have evolved various life-history strategies to survive fluctuating resource conditions in soils. However, it remains elusive how the life-history strategies of microorganisms influence their processing of organic carbon, which may affect microbial interactions and carbon cycling in soils. Here, ENIGMA researchers characterized the genomic traits, exometabolite profiles, and interactions of soil bacteria representing copiotrophic and oligotrophic strategists. I
Two model phages characterized by new CRISPR based technology extensible to diverse phages
ENIGMA researchers demonstrated for the first time that they can, on a genome-wide scale, identify phage genes that are essential (or not) to infecting bacteria, and then replace non-essential DNA with distinctive barcode tags. Their method could unlock potent biotechnology applications.
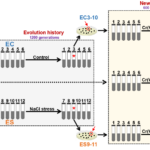
Environmental bacteria may have candidate genes for bioremediation
ENIGMA researchers identified important genes for improving Chromate resistance in genetically diverse bacteria through experimental evolution.
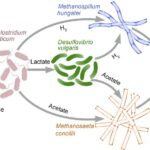
To Study Competition and Cross-Feeding, Scientists Build Synthetic Microbiomes
A new study investigated complex interactions among four cross-feeding microorganisms in a synthetic community (SynCom) that converts cellulose to methane and carbon dioxide, paving the way toward a more predictive understanding of the impact of environmental perturbations on microbial interactions sustaining geochemically significant processes in natural systems.