A collaboration of researchers at Johns Hopkins University, Colorado State University, The University of Tennessee, The Lawrence Berkley National Laboratory, The Massachusetts Institute of Technology, The University of Oklahoma and the University of Georgia, Oak Ridge National Laboratory researchers in the Bioscience Division for the ENIGMA have released an article in Environmental Science and Technology.
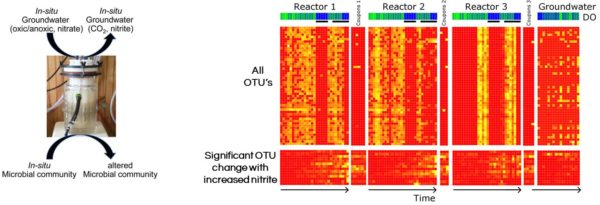
They have developed a novel in-field bioreactor system for closely approximating in-situ conditions and monitoring the response of the groundwater microbial communities to geochemical manipulation beyond the bulk phase of traditional systems. The effect of contaminants on the diversity, structure, function, and biotransformation capabilities of groundwater microbial communities have been studied at the Oak Ridge Field Research Center (ORFRC), but only provided limited data linking geochemistry to microbial community structure and ecological function. “The advantage of this new system is that these bioreactors are much smaller and can be mobile, which allows for greater experimental replication and flexibility. We have also incorporated both the free-living or planktonic organisms as well as those in biofilms in the same vessel so we can monitor both simultaneously” said Dwayne Elias the lead researcher on the study. Due to the constant influx of native groundwater, in situ conditions can be maintained for considerable lengths of time as opposed to closed microcosms. “Not only can we monitor the site conditions for long periods, but we can also introduce perturbations to some of the bioreactors and observe the community response as compared to the ongoing, actual site condition bioreactors, which we hope can inform site and risk management decisions in the future”, Elias said.
Significance
- Successfully monitored microbial community for >16 weeks with reproducible community responses to repeated perturbations.
- Provides evidence of stochastic microbial community assembly.
- Ties the lab to the field by providing a technology for testing lab-based hypotheses in the field.
