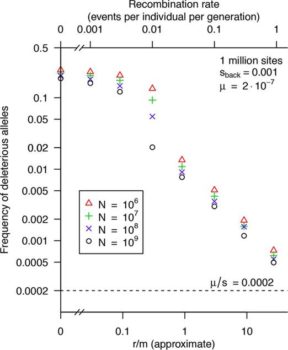
Free-living bacteria are usually thought to have large effective population sizes, and so tiny selective differences can drive their evolution. However, because recombination is infrequent, “background selection” against slightly deleterious alleles should reduce the effective population size (Ne) by orders of magnitude. For example, for a well-mixed population with 1012 individuals and a typical level of homologous recombination (r/m = 3, i.e., nucleotide changes due to recombination [r] occur at 3 times the mutation rate [m]), we predict that Ne is <107. An argument for high Ne values for bacteria has been the high genetic diversity within many bacterial “species,” but this diversity may be due to population structure: diversity across subpopulations can be far higher than diversity within a subpopulation, which makes it difficult to estimate Ne correctly. Given an estimate of Ne, standard population genetics models imply that selection should be sufficient to drive evolution if Ne × s is >1, where s is the selection coefficient. We found that this remains approximately correct if background selection is occurring or when population structure is present. Overall, we predict that even for free-living bacteria with enormous populations, natural selection is only a significant force if s is above 10e7 or so.
Price, MN, Arkin, AP. Weakly deleterious mutations and low rates of recombination limit the impact of natural selection on bacterial genomes (2015)mBio, doi: 10.1128/mBio.01302-15
