Microbiology studies benefit immensely from the ability to genetically manipulate the bacterial strain of interest. Transition from purely genomic studies to genetics-enabled molecular biology approaches, often relies on plasmids, or small circular DNA elements distinct from the chromosome. In developing plasmid-based systems, knowledge of the origin of replication (the point at which DNA replication begins) that works with the bacteria’s DNA maintenance machinery, is needed. However, many newly discovered bacterial strains lack known plasmids or characterized origins of replication.
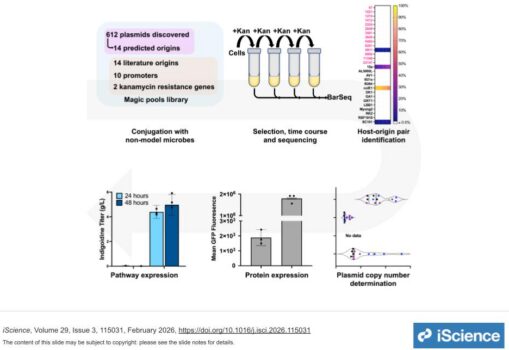
Codik et al. puts forward a way to validate predicted origins of replication in newly discovered or existing model bacteria. This work led to the discovery of three previously unknown origins of replication, one of which was functional in Pantoea MT58. To validate this newly discovered origin of replication in Pantoea, the bacterium was engineered to produce non-native natural products such as a non-ribosomal peptide compound and a terpenoid. This work opens new doors for biosystems design and improves our understanding of bacterial functions.
Impact to Field/Science
Plasmids are important extrachromosomal and often mobile genetic elements that encode functions critical to microbial ecosystems and functions. They also inherently contain biological parts necessary to develop genetic tools in new environmental isolates to facilitate their study. The results in this study leverage previously identified plasmids and their origins of replication to bring genetic engineering to non-model bacterial systems. Introduction of new origins of replication in our toolkit allows deeper understanding into how these microbial systems function as well as what they can be used for.
Significance to ENIGMA
ENIGMA routinely isolates new bacterial strains but can only mount deep molecular biology studies in a subset where genetic manipulation is possible. Prior discovery projects in ENIGMA led to the discovery of very large numbers on new plasmids in ground water samples. Our previous legacy ENIGMA study provided a compelling resource. Used along with publicly available origin sequence prediction tools, the current study resulted in a valuable approach to discover and validate new origins of replication in environmentally important bacteria, isolated in the ENIGMA project. These isolates can now be genetically manipulated for deeper characterization of how they can survive under such harsh conditions at the Oak Ridge Reservation (ORR).
Collaborations
Alex Codik1,2, Ankita Kothari1, Hualan Liu3, Benjamin L. Weinberg4, Trenton K. Owens4, Aparajitha Srinivasan1, Alex Rivier1, Thomas Eng1, Adam P. Arkin2,4,6, Adam M. Deutschbauer4,5, Aindrila Mukhopadhyay1,2*
- Biological Systems and Engineering Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA
- Comparative Biochemistry Graduate Group, University of California, Berkeley, Berkeley, CA 94720, USA
- US Department of Energy Joint Genome Institute, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA
- Environmental Genomics and Systems Biology Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA
- Department of Plant and Microbial Biology, University of California, Berkeley, CA 94720, USA
- Department of Bioengineering, University of California, Berkeley, CA 94720, USA
