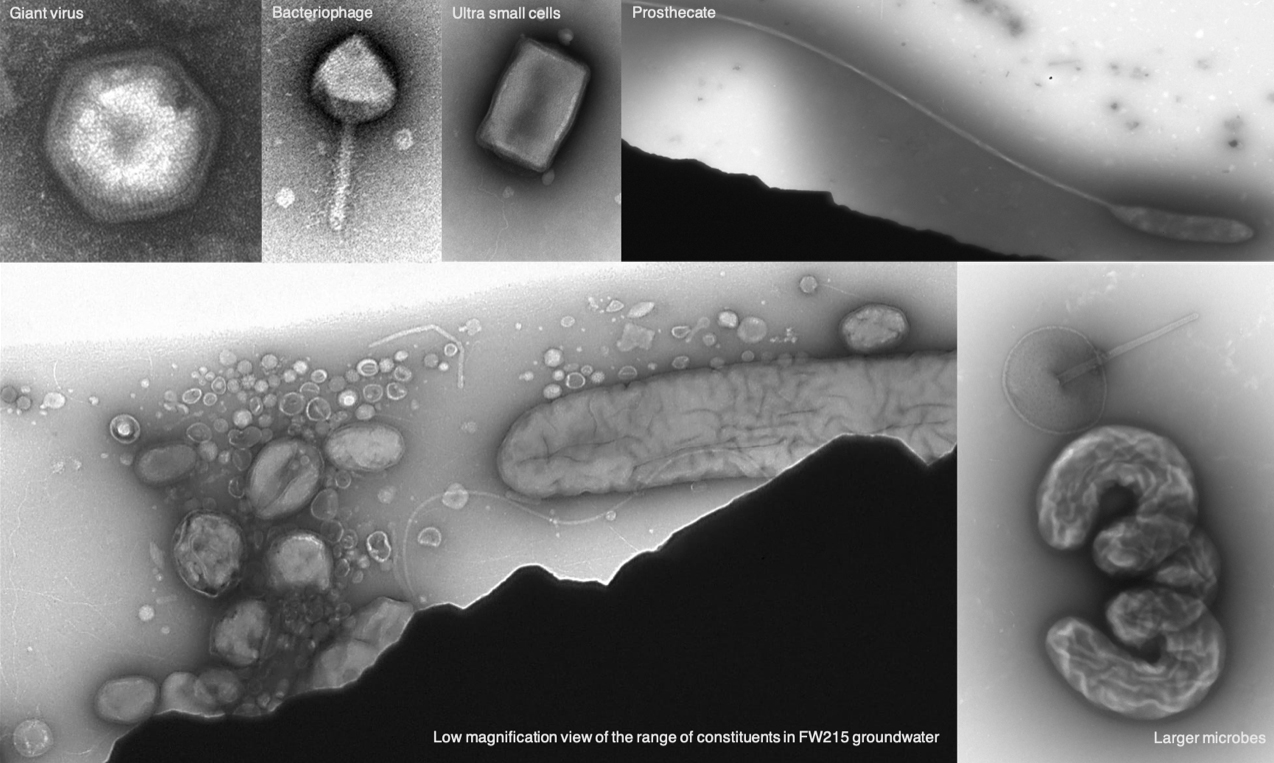
Lab-based ENIGMA researchers enrich, isolate and physiologically/genetically characterize the specific taxa that are predicted to be the most important to the observed field processes. By characterizing genetic functions and linking genotype to phenotype in the laboratory, we can interpret field data mechanistically, leading to pointed hypotheses of the role of particular taxa, proteins, co-factors, and their interactions in creating systems behavior.

Relevant Publications
 New Origins of Replication Expand Our Toolkit for Understanding and Engineering Microbial Systems - Microbiology studies benefit immensely from the ability to genetically manipulate the bacterial strain of interest. Transition from purely genomic studies to genetics-enabled molecular biology approaches, often relies on plasmids, or small circular DNA elements distinct from the chromosome. In developing plasmid-based systems, knowledge of the origin of replication (the point at which DNA replication begins) that works with the bacteria’s… More →
New Origins of Replication Expand Our Toolkit for Understanding and Engineering Microbial Systems - Microbiology studies benefit immensely from the ability to genetically manipulate the bacterial strain of interest. Transition from purely genomic studies to genetics-enabled molecular biology approaches, often relies on plasmids, or small circular DNA elements distinct from the chromosome. In developing plasmid-based systems, knowledge of the origin of replication (the point at which DNA replication begins) that works with the bacteria’s… More →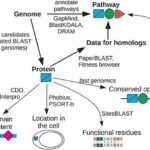 Interactive tools for functional annotation of bacterial genomes - ###How to predict a protein’s function or a bacterium’s capabilities Predicting what a bacterium can do from its genome is a complex task that involves understanding the functions of its encoded proteins. While automated software for predicting protein function is widely used, it is prone to errors, with over 10% of automated annotations for bacterial enzymes and transporters being incorrect.… More →
Interactive tools for functional annotation of bacterial genomes - ###How to predict a protein’s function or a bacterium’s capabilities Predicting what a bacterium can do from its genome is a complex task that involves understanding the functions of its encoded proteins. While automated software for predicting protein function is widely used, it is prone to errors, with over 10% of automated annotations for bacterial enzymes and transporters being incorrect.… More → New Tool Enables Overexpression Libraries for High-throughput Protein Characterization - The development of a high-throughput method by for determining the function of predicted proteins of unknown function marks a significant advancement in the field of microbial biology. By utilizing DNA barcode sequencing and auxotrophic strains of E. coli, ENIGMA Researchers at Lawrence Berkeley National Laboratory can now assign functions to many previously uncharacterized genes. This breakthrough enables more complete identification… More →
New Tool Enables Overexpression Libraries for High-throughput Protein Characterization - The development of a high-throughput method by for determining the function of predicted proteins of unknown function marks a significant advancement in the field of microbial biology. By utilizing DNA barcode sequencing and auxotrophic strains of E. coli, ENIGMA Researchers at Lawrence Berkeley National Laboratory can now assign functions to many previously uncharacterized genes. This breakthrough enables more complete identification… More → Microbial Remains Shape Community Composition and Growth of Soil Microorganisms - The Science Microbes, which include organisms like bacteria, are tiny living things that inhabit virtually every environment on Earth. Soil microbes play a crucial role in breaking down organic matter and are crucial drivers of global nutrient cycles that support all terrestrial ecosystems. ENIGMA scientists studied what happens when these microorganisms die and leave behind remains known as necromass. After… More →
Microbial Remains Shape Community Composition and Growth of Soil Microorganisms - The Science Microbes, which include organisms like bacteria, are tiny living things that inhabit virtually every environment on Earth. Soil microbes play a crucial role in breaking down organic matter and are crucial drivers of global nutrient cycles that support all terrestrial ecosystems. ENIGMA scientists studied what happens when these microorganisms die and leave behind remains known as necromass. After… More → Goff Uses KBase to Analyze Genetic Factors Impacting Microbial Fitness in Contaminated Environments - A recent KBase community highlight summarized research accomplishments from Jennifer Goff, who was a postdoctoral researcher with the ENIGMA Science Focus Area in Mike Adams' lab at the University of Georgia (UGA) and is now a professor at SUNY College of Environmental Science and Forestry (ESF). Several of her team’s latest studies, which used KBase to conduct large-scale genetic analyses,… More →
Goff Uses KBase to Analyze Genetic Factors Impacting Microbial Fitness in Contaminated Environments - A recent KBase community highlight summarized research accomplishments from Jennifer Goff, who was a postdoctoral researcher with the ENIGMA Science Focus Area in Mike Adams' lab at the University of Georgia (UGA) and is now a professor at SUNY College of Environmental Science and Forestry (ESF). Several of her team’s latest studies, which used KBase to conduct large-scale genetic analyses,… More → Machine Learning Reveals Patterns of REC Protein Domain Evolution - Bacteria can evolve by exchanging genetic material through a process called recombination. This study used machine learning models combined with functional laboratory experiments to show that after recombination, a bacterial signaling system changed and expanded its protein family, despite considerable selective pressures in place that constrained new modifications. More →
Machine Learning Reveals Patterns of REC Protein Domain Evolution - Bacteria can evolve by exchanging genetic material through a process called recombination. This study used machine learning models combined with functional laboratory experiments to show that after recombination, a bacterial signaling system changed and expanded its protein family, despite considerable selective pressures in place that constrained new modifications. More →