 ENIGMA’s Vivek Mutalik, Adam Deutschbauer, Pavel Novichkov, and Adam Arkin win R&D100 Award for a tool developed under our “Discovery Program”; where high-risk projects are funded for a short duration to encourage high impact changes in science or technological capability that extend and enhance our ongoing research
ENIGMA’s Vivek Mutalik, Adam Deutschbauer, Pavel Novichkov, and Adam Arkin win R&D100 Award for a tool developed under our “Discovery Program”; where high-risk projects are funded for a short duration to encourage high impact changes in science or technological capability that extend and enhance our ongoing research
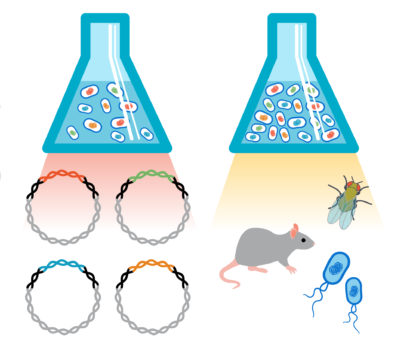
Dub-seq technology is based on creating a genomic fragment library in association with dual barcodes on broad-host vectors. DNA (from any source) is sheared and cloned between dual barcodes. Then, the cloned genomic fragment is identified and associated with unique DNA barcode upfront and is done only once for each library. Once this step of associating barcodes to genomic fragment is performed, only one of the barcodes is used as a proxy for clonal isolate in the following experiments. This standardization enables researchers to assay the same library across diverse conditions with minimal cost per genome-wide assay. A simple analytic approach aids in connecting fragment score and gene score to gene function.
This technology fills a critical gap in elucidating gene-function in a very high-throughput assay set up and saves time, labor, and money as compared to state-of-the-art methods. Dub-seq technology is applicable in the discovery of novel enzymes/biocatalysts for biofuel production, improving and tolerance traits for toxic biochemicals, ensemble functional assessment of microbial communities, plant growth-promoting factors, bioremediation routes, novel green chemistries and biotechnologies in improving energy and environment missions. The technology is also extendable in diverse health and agriculture associated biotechnologies.
For more information associated with the award: https://newscenter.lbl.gov/2017/11/20/berkeley-lab-takes-home-four-rd-100-awards-for-cutting-edge-technologies/

