Using microcalorimetry and thermodynamic modeling to examine effects of thermal stress
Hunt KA, von Netzer F, Gorman-Lewis D, Stahl DA. (2022) Microbial maintenance energy quantified and modeled with microcalorimetry. Biotechnol Bioeng. 2022 Jun 9; [DOI]:10.1002/bit.28155. {PMID}: 35680566 OSTI:1874049 All known biology is driven by the partitioning of energy released from nutrient consumption, either into building new cells or maintaining cellular integrity required for viability. The energy […]
Discovering microbial adaptations to subsurface niches
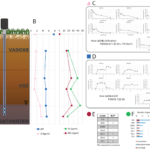
Gushgari-Doyle, S; L.M. Lui, T.N. Nielsen, X. Wu, R.G. Malana, F.L. Poole II, M.W.W. Adams, A.P. Arkin, R. Chakraborty (2022) Genotype to ecotype in niche environments: Adaptation of Arthrobacter to carbon availability and environmental conditions. ISME Comm. [DOI]:10.1038/s43705-022-00113-8 OSTI:1860268 Different niche micro-environments can affect the structure and function of microbial communities. Terrestrial subsurface environments exhibit […]
Sampling Down Well In Situ Microbial Colonization Conditions
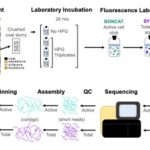
LJ McKay*, HJ Smith*, E Barnhart, HD Schweitzer, RR Malmstrom, D Goudeau, MW Fields (2021) Activity-based, genome-resolved metagenomics uncovers key populations and pathways involved in subsurface conversions of coal to methane Nature ISME Journal [DOI]:10.1038/s41396-021-01139-x PMID: 34689183 Development of activity-based metagenomics, enabled the identification of translationally active microorganisms and associated functional capacity Demonstration on terrestrial subsurface […]
Uncovering metal tolerance mechanisms using ENIGMA tools and Analysis
Thorgersen, M.P., .; J. Xue, E.L.W. Majumder, V.V. Trotter, X. Ge, F.L. Poole II, T.K. Owens, L.M. Lui, T.N. Nielsen, A.P. Arkin, A.M. Deutschbauer, G. Siuzdak, and M.W.W. Adams Deciphering microbial metal toxicity responses using RB-TnSeq and activity-based metabolomics. Applied and Environmental Microbiology 87 (21), e01037-21 (2021). [DOI]: 10.1128/AEM.01037-21] {PMID}:34432491 OSTI:1825256 Deciphering Microbial Metal […]
Semi-Automated Complete Microbial Genome Assembly
Lui, Lauren M.; T.N. Nielsen, A.P. Arkin (2021) A method for achieving complete microbial genomes and better-quality bins from metagenomics data. PloS Computational Biology [DOI]:1371/journal.pcbi.1008972 OSTI:1788019 A Method for Achieving Complete Microbial Genomes and Improving Bins From Metagenomics Data Assembling complete microbial genomes from short read metagenomics data is difficult, but the method “Jorg” helps […]
Observing the Microbial Genome
Basics2Breakthroughs: Observing the microbial genome from Berkeley Lab on YouTube. ENIGMA Project Scientist Lauren Lui is profiled on LBL Basics2Breakthroughs, filmed for LBNL’s 90th anniversary. She uses the latest bioinformatics technology to study microbes so we can learn how their genes shape the environment. Her ultimate goal is to be able to predict microbial interactions […]
Clarivate Globally Highly Cited Researcher Jizhong Zhou
ENIGMA’s Jizhong Zhou from University of Oklahoma, Norman, is once again listed by Clarivate as a Globally Highly Cited Researcher for 2021; judged by the numbers of publications which are 1% highly cited. Globally less than 6,400 researchers are recognized based on the information from the database. A selection of his highly cited papers supported […]