Using new barcoded screening technology, ENIGMA researchers uncovered genome-wide targets of diverse including single-stranded RNA-phages (ssRNA-phages)-derived protein antibiotics to efficiently identify the host mechanisms targeted in bacteriolysis. The novel approach could have significant applications for characterizing the hundreds of putative Sgl identified in genomic databases and recent studies.
ENIGMA researchers awarded for First Place Talks
University of Georgia ENIGMA researchers Jennifer Goff (postdoctoral fellow) and Elizabeth Szink (undergraduate research associate) presented their work at the 10th Annual Southeastern Biogeochemistry Symposium May 12-14,2023 at the University of South Carolina, Columbia, SC . The Southeastern Biogeochemistry Symposium is a unique opportunity for undergraduate students, graduate students, post doctorate, and faculty members from the […]
ENIGMA researchers Win 2nd Annual BioEPIC SLAM
Adam Arkin, lead scientist, along with ENIGMA team members Bradley Biggs , a postdoctoral researcher in the Arkin lab, and Mingfei Chen, a postdoctoral researcher in the Chakraborty lab, recently participated in a virtual Research Slam promoting science related to the BioEPIC building. https://bioepic.lbl.gov/slam/ Results: Mingfei Chen (ENIGMA) took first place, Mo Ombadi (Watershed […]
ENIGMA scientists highly cited across multiple platforms
ENIGMA scientists Paul Adams and Jizhong Joe Zhou are included in Clarivate’s Web of Science Highly Cited Researchers who have demonstrated significant and broad influence reflected in their publication of multiple highly cited papers over the last decade.These highly cited papers rank in the top 1% by citations for a field or fields and publication […]
Jizhong Zhou elected to the American Academy of Arts and Sciences
Jizhong Joe Zhou (ENIGMA principal investigator, affiliate faculty in Berkeley Lab’s Earth & Environmental Sciences Area, director of the Institute for Environmental Genomics at the University of Oklahoma, has been elected as a member of American Academy of Arts and Science (AmAcad). He will participate, along with the other newly elected members, in the 2023 […]
Examining adaptations of a dominant bacterial species at a contaminated subsurface site
Leveraging comparative genomics, community analysis, and geochemical data to understand adaptations that facilitate survival of a dominant community member in the contaminated Oak Ridge Reservation subsurface.
ENIGMA researchers identify hundreds of new enzymes
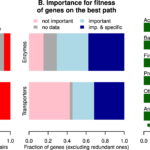
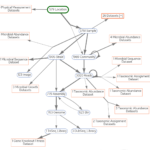
Filling Gaps in Bacterial Metabolism with Computation and High-throughput Genetics: An analysis of genetic data from many different kinds of bacteria identified hundreds of new enzymes.
The Contextual Ontology-based Repository Analysis Library (CORAL) facilitates FAIR ENIGMA datasets
Novichkov, PS*; Chandonia, J-M*; Arkin, AP. (2022) CORAL: A framework for rigorous self-validated data modeling and integrative, reproducible data analysis. GigaScience [DOI]:10.1093/gigascience/giac089 OSTI ID: 1888047 The Contextual Ontology-based Repository Analysis Library (CORAL) platform greatly facilitates adherence to the FAIR principles, including the especially difficult challenge of making heterogeneous datasets Interoperable and Reusable across all […]